Some information about Scholia
Scholia is mostly a web service developed from GitHub at https://github.com/fnielsen/scholia in an open source fashion. It was inspired by discussions at the WikiCite 2016 meeting in Berlin. Anyone can contribute as long as their contribution is under GPL.
I started to write the Scholia code back in October 2016 according to the initial commit at https://github.com/fnielsen/scholia/commit/484104fdf60e4d8384b9816500f2826dbfe064ce Since then particularly Daniel Mietchen and Egon Willighagen have joined in and Egon has lately be quite active.
Users can download the code and run the web service from their own computer if they have a Python Flask development environment. Otherwise the canonical web site for Scholia is https://tools.wmflabs.org/scholia/ which anyone with an Internet connection should be able to view.
So what does Scholia do? The initial “application” was a “static” web page with a researcher profile/CV of myself based on data extracted from Wikidata. It is still available from: http://people.compute.dtu.dk/faan/fnielsenwikidata.html. I added a static page for my research section, DTU Cognitive Systems, showing scientific page production and a coauthor graph. This is available here: http://people.compute.dtu.dk/faan/cognitivesystemswikidata.html.
The Scholia web application was an extension of these initial static pages so a profile page for any researcher or any organization could be made on the fly. And now it is no longer just authors and organizations where there is a profile page, but also works, venues (journals or proceedings), series, publishers, sponsors (funders) and awards. We have also “topics” and individual pages showing specialized information about chemicals, proteins, diseases and biological pathways. A rudimentary search interface is implemented.
The content of the web pages of Scholia with plots and tables are made from queries to the Wikidata Query Service, – the extended SPARQL endpoint provided by the Wikimedia Foundation. We also pull in text from the introduction of the articles in the English Wikipedia. We modify the table output of the Wikidata Query Service so individual items displayed in table cells link back to other items in Scholia.
Egon Willighagen, Daniel Mietchen and I have described Scholia and Wikidata for scientometrics in the 16-pages workshop paper “Scholia and scientometrics with Wikidata” https://arxiv.org/pdf/1703.04222.pdf The screenshots shown in the paper has been uploaded to Wikimedia Commons. These and other Scholia media files are available in category page https://commons.wikimedia.org/wiki/Category:Scholia
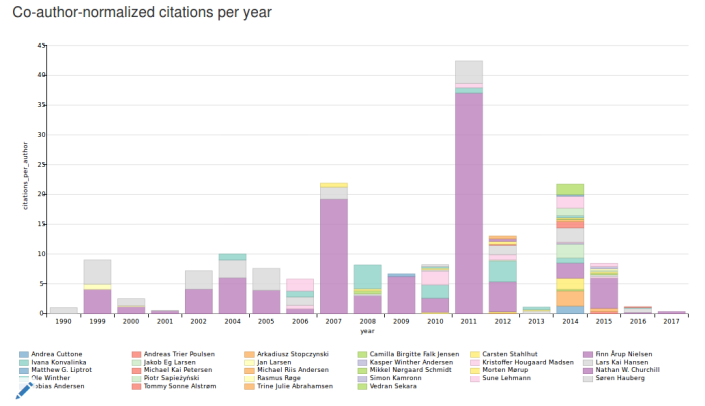
Working with Scholia has been a great way to explore what is possible with SPARQL and Wikidata. One plot that I like is the “Co-author-normalized citations per year” plot on the organization pages. There is an example on this page: https://tools.wmflabs.org/scholia/organization/Q24283660. Here the citations to works authored by authors affiliated with the organization in question are counted and organized in a colored bar chart with respect to year of publication, – and normalized for the number of coauthors. The colored bar charts have been inspired by the “LEGOLAS” plots of Shubhanshu Mishra and Vetle Torvik.
Part of the Python Scholia code will also work as a command-line script for reference management in the LaTeX/BIBTeX environment using Wikidata as the backend. I have used this Scholia scheme for a couple of scientific papers I have written in 2017. The particular script is currently not well developed, so users would need to be indulgent.
Scholia relies on users adding bibliographic data to Wikidata. Tools from Magnus Manske are a great help as are Fatameh of “T Arrow” and “Tobias1984” and the WikidataIntegrator of the GeneWiki people. Daniel Mietchen, James Hare and a user called “GZWDer” have been very active adding much of the science bibligraphic information and we are now past 2.3 million scientific articles on Wikidata. You can count them with this link: https://tinyurl.com/yaux3uac